We provide online services for non-commercial use, powered by AmoAi, which include innovative tools such as Evoid®, Eimo®, RSign®, PhiSign®, PhiGnet®, AmoAi®, Direct3D®, RamaPot®, DaVis®, Phimo®, and PAD®. By accessing and using these services and the accompanying software provided by AmoAi ("Amo" or "we"), you agree to abide by the terms outlined in the License Agreement between you and AmoAi.
Infer disease variants for clinical decisions and therapeutics
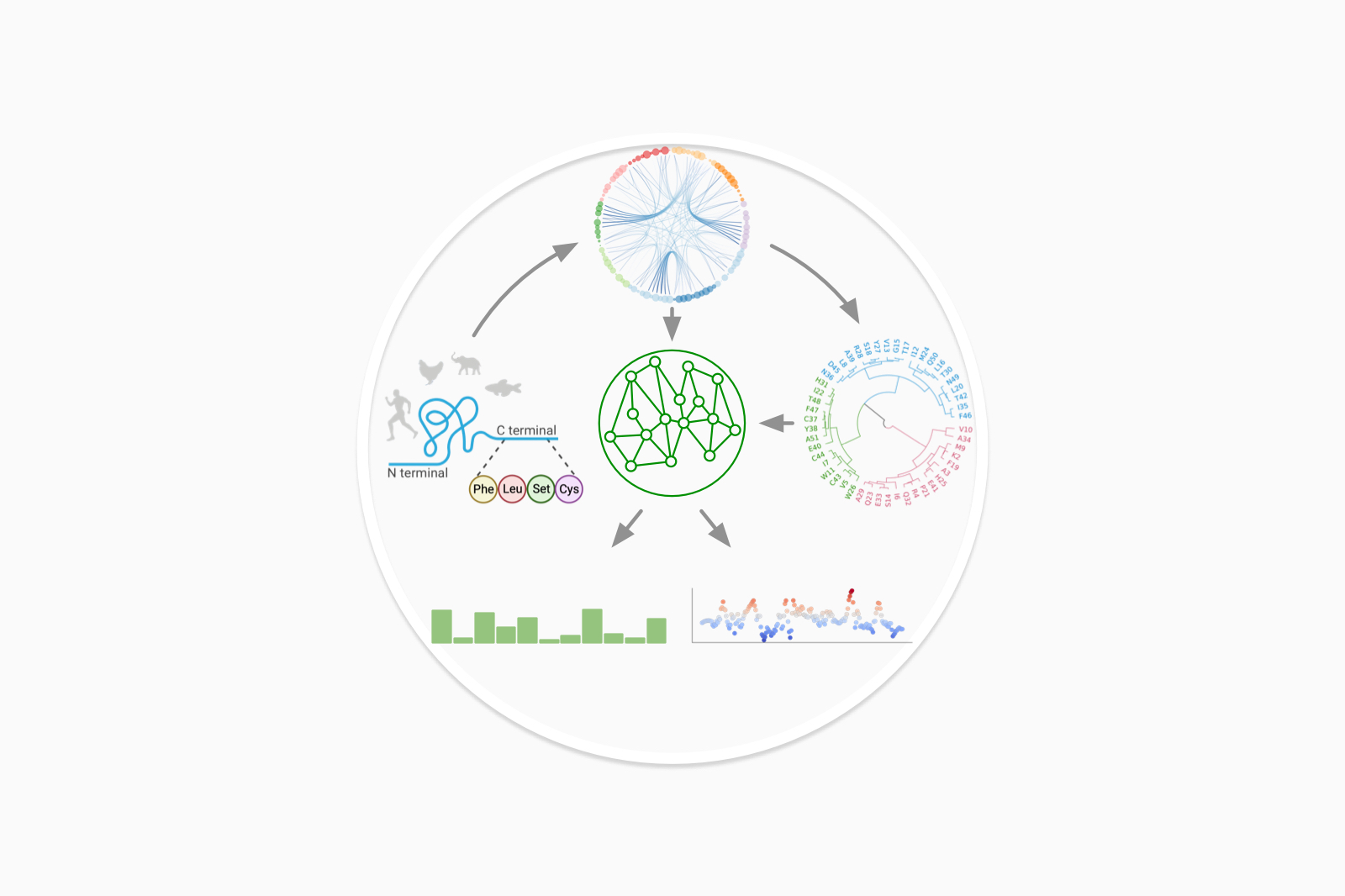
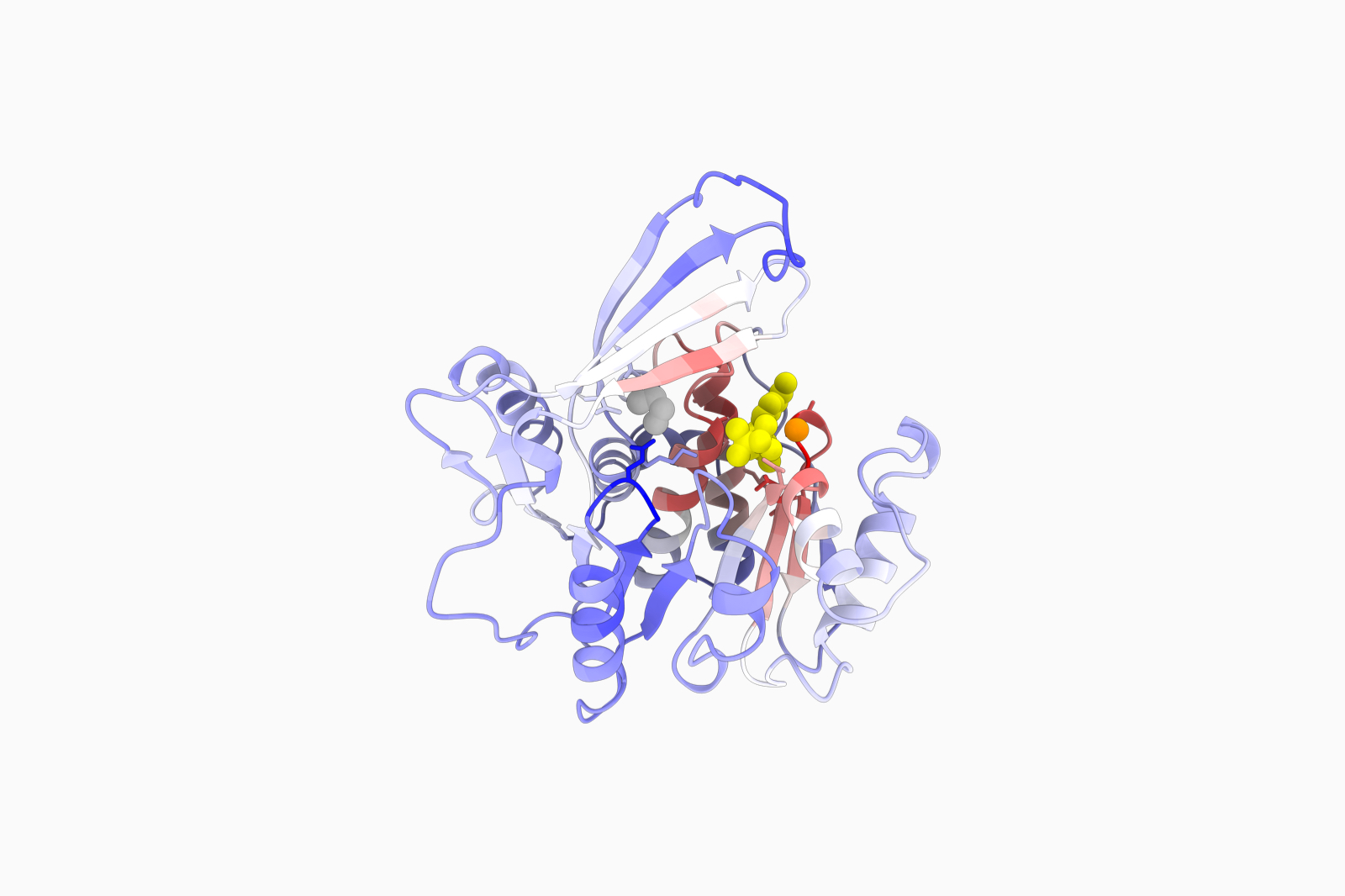
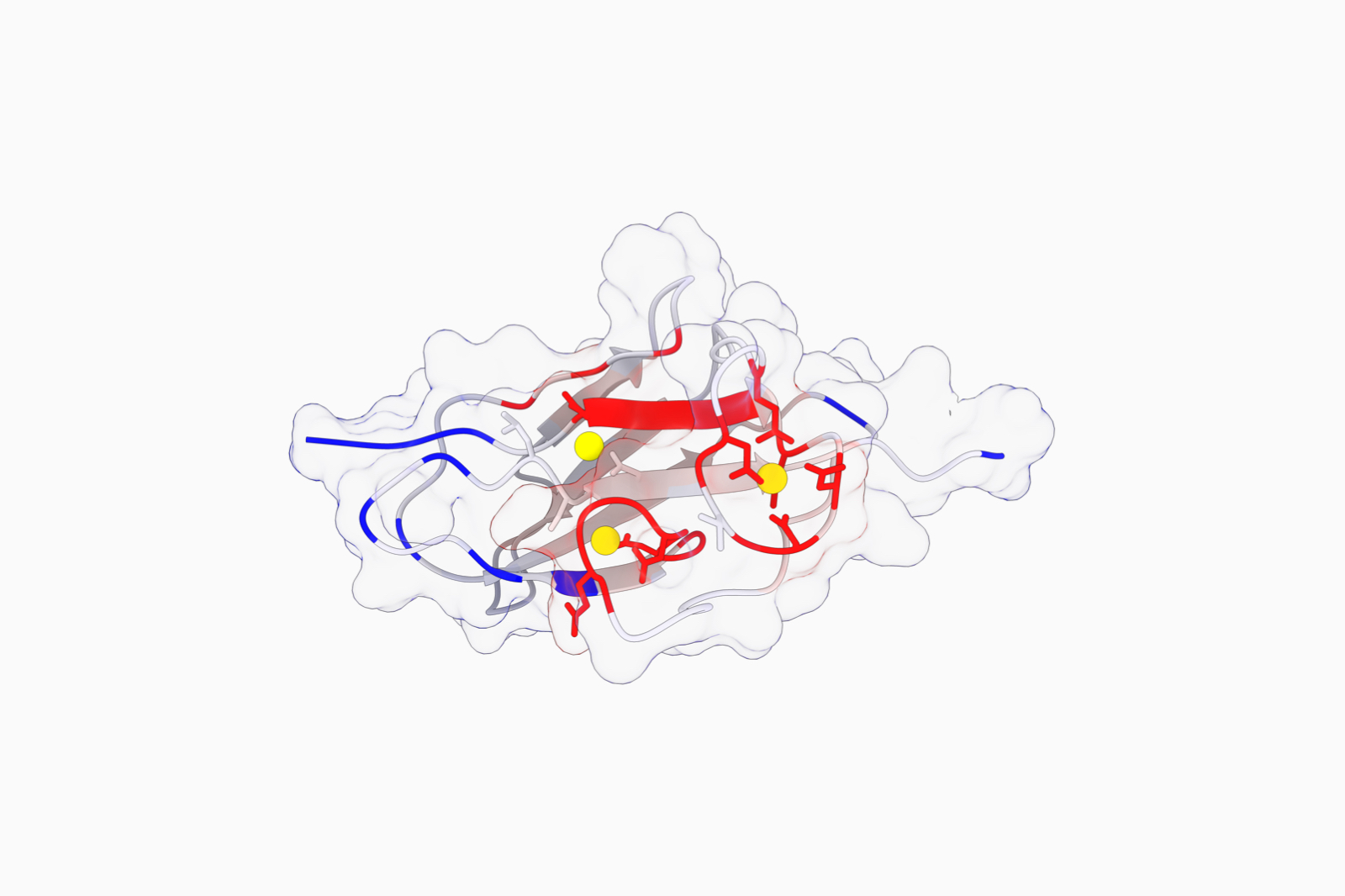
Infer protein function at the residue level from its sequence
Analyze biological sequences using bioinformatic methods
Design proteins of desired function via generative learning
Design proteins of desired function using machine learning
Infer protein function using evolutionary data
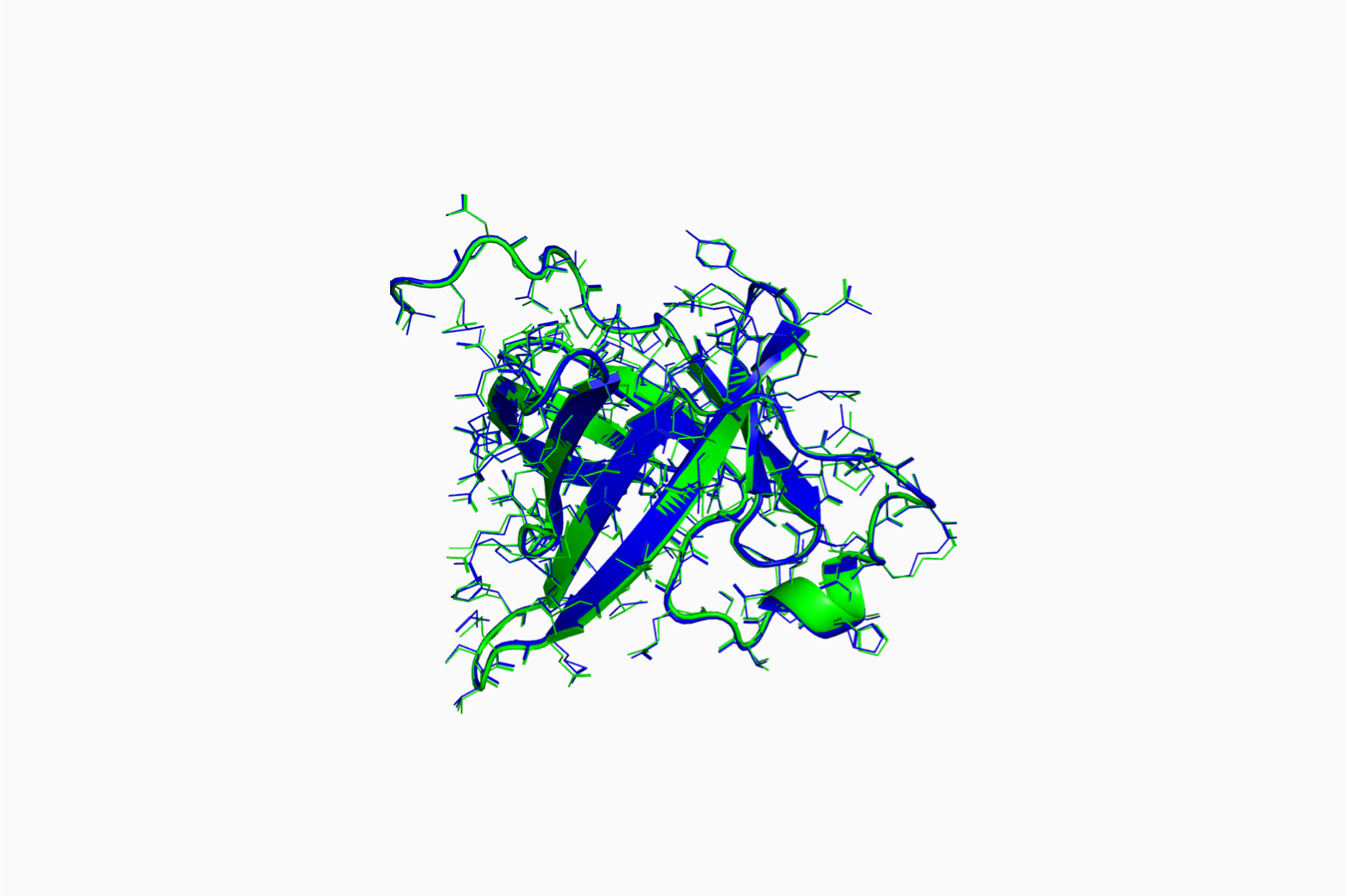
Predict protein 3D structures using deep learning
Devise cutting-edge machine learning approaches
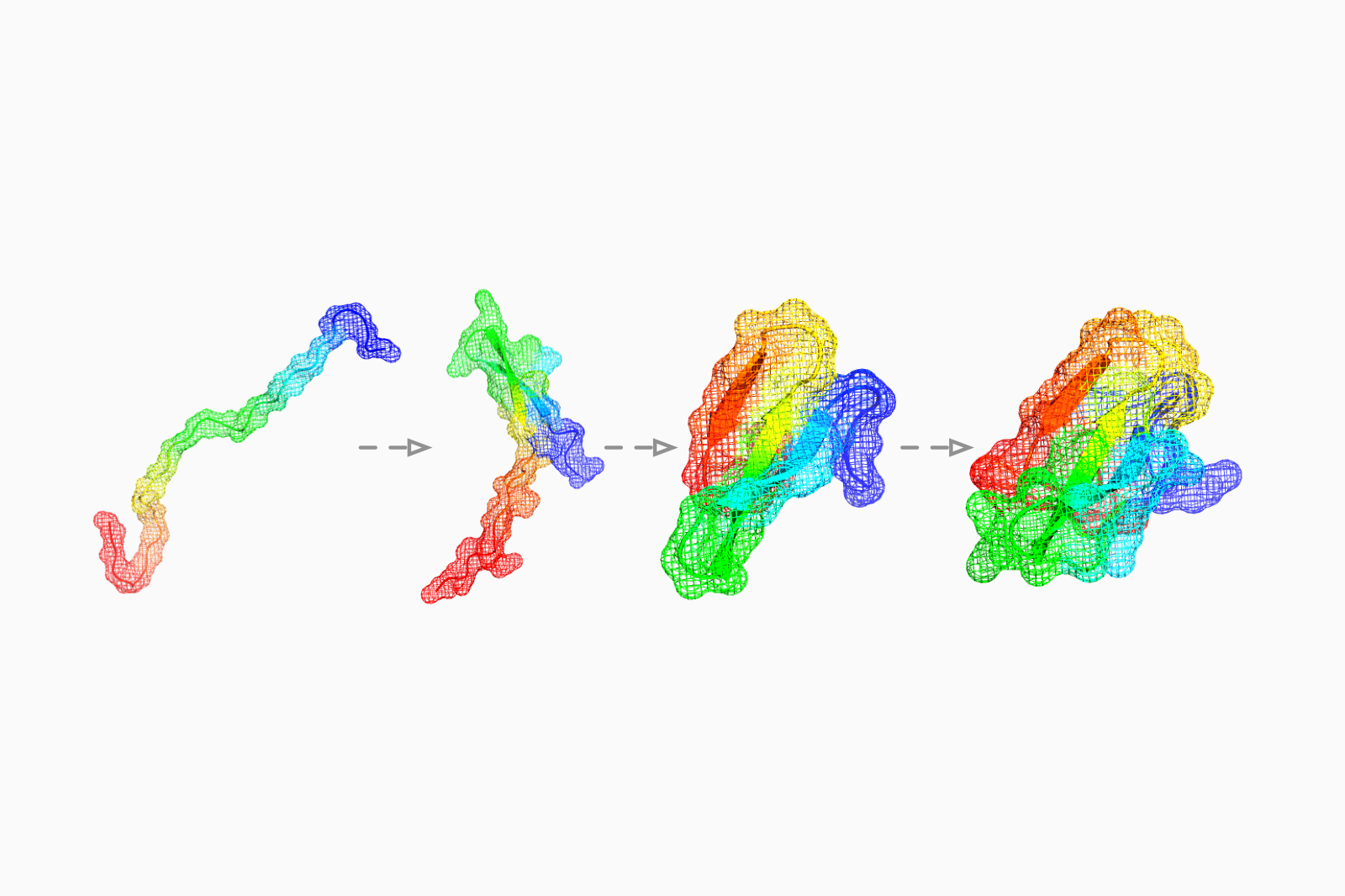
Predict folding pathways using graph networks
Compute Ramachandran basins for protein sequences
Visualize high-dimensional data interactively in 2/3D space
Provide large-scale datasets for machine learning
Evoid®
Version 1.0.0
Webserver
Data sets
Tutorial
Last update: 2023-9-15
Eimo®
Version 2.0.0
Webserver
Data sets
Tutorial
Last update: 2023-9-15
AmoAi® tools
Version 2.8.0
Webserver
Data sets
Tutorial
Last update: 2023-9-15
PhiGnet®
Version 1.0.9
Webserver
Data sets
Tutorial
Last update: 2023-9-15
Direct3D®
Version 1.0.0
Webserver
Data sets
Tutorial
Last update: 2023-9-15
DaVis®
Version 1.2.0
Webserver
Data sets
Tutorial
Last update: 2023-9-15