Protein-based therapeutics
How to develop the next generation therapeutics?
G protein-coupled receptors (GPCRs) stand as a pivotal and extensive integral membrane protein family, holding paramount physiological significance. Their responsiveness to diverse physiological and environmental stimuli makes them crucial players in cellular signaling. Remarkably, GPCRs are key therapeutic targets for a myriad of human diseases, constituting approximately one-third of the drugs currently available in the market. Recent breakthroughs in GPCR structural biology have revolutionized our comprehension of GPCR function, ligand binding, and pharmacological actions. This progress not only enhances our understanding of these receptors but also propels advancements in drug design.
Single mutagenesis
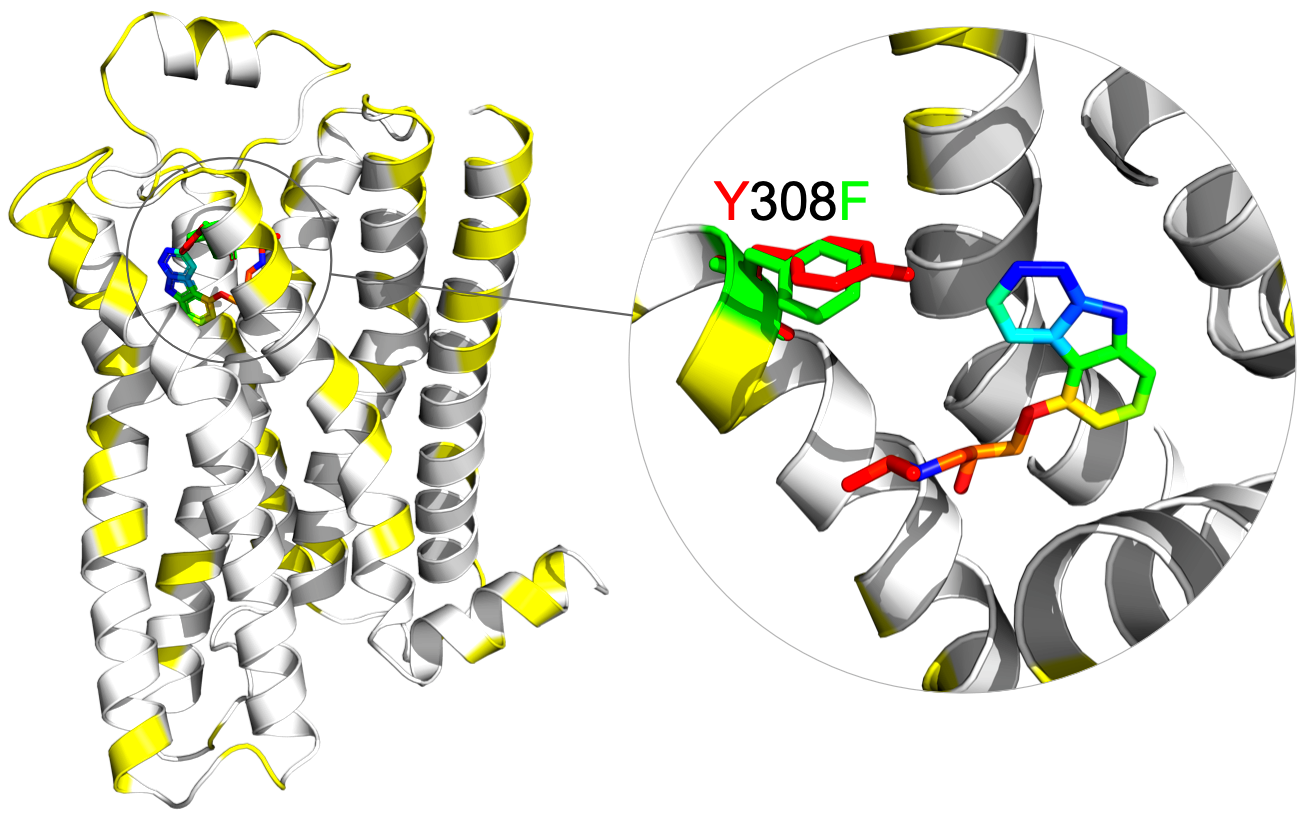
Where are coupled residues located?
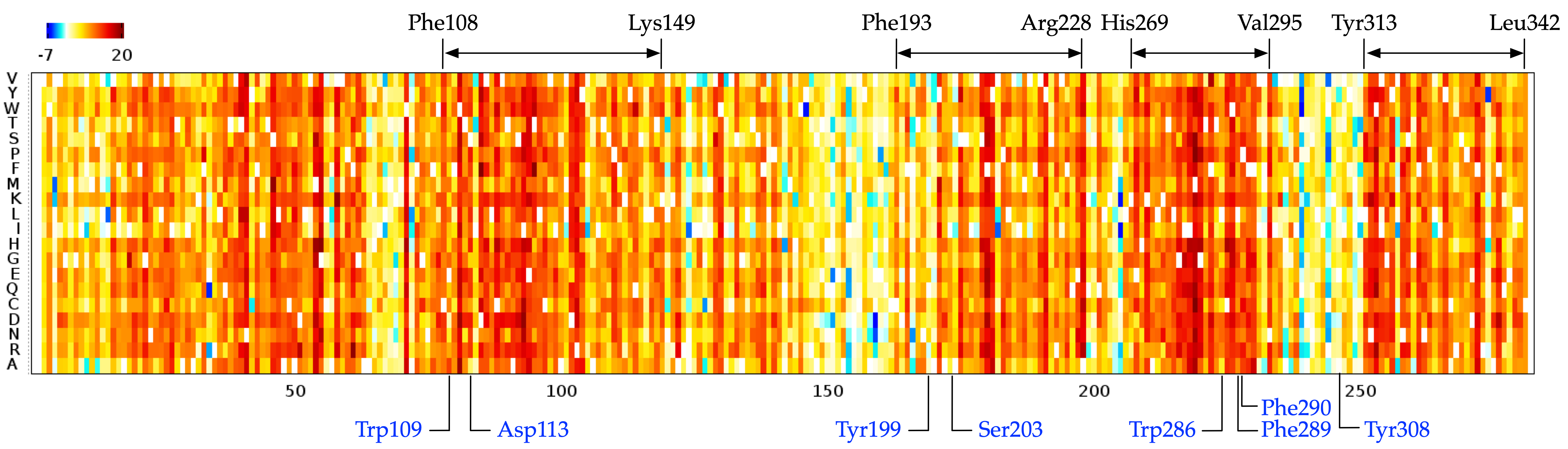
Complete single mutagenesis. The matrix that is computed by the SAEC method shows \(\Delta E\) — the energy difference of each mutant sequence with each mutation \(\tau\) at the ith site and wild-type (WT) sequence, negative values representing favourable while positive representing unfavourable mutations.
Sequence design
Where are coupled residues located?
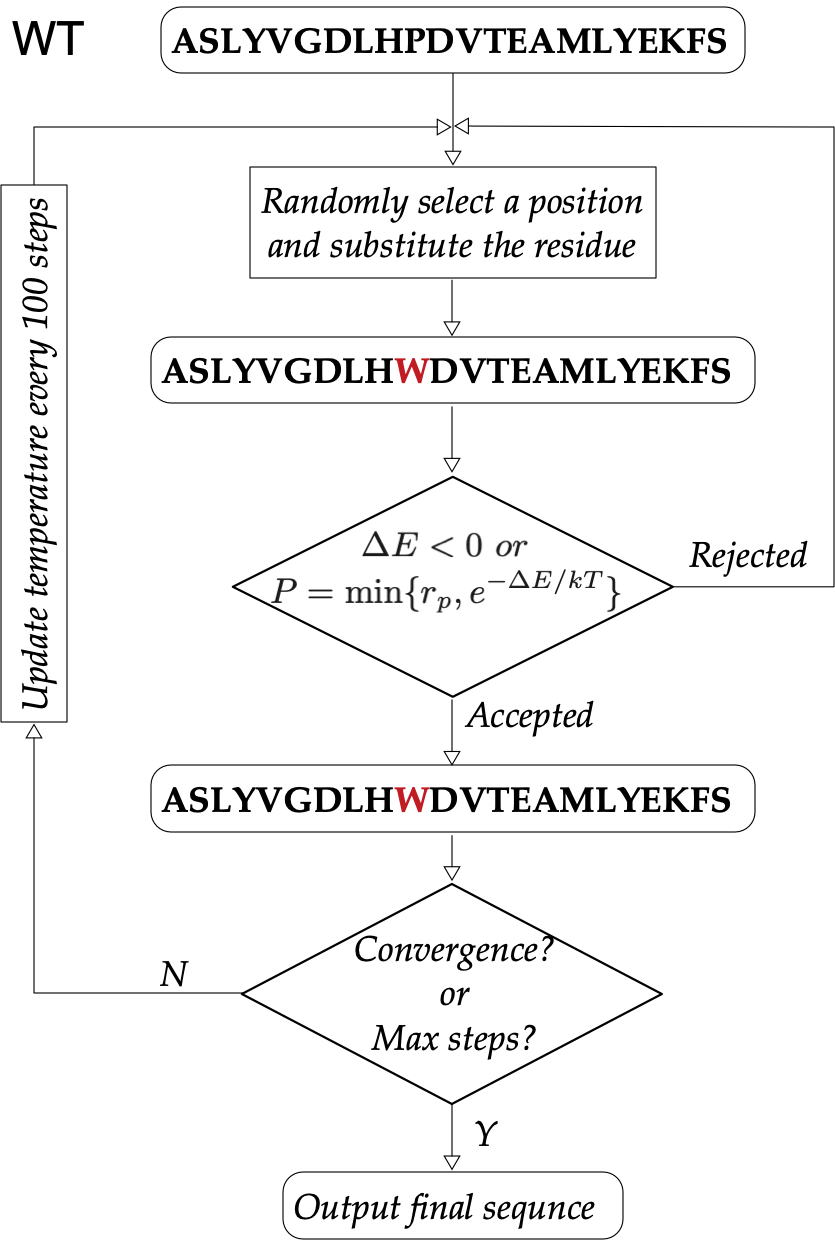
Starting from a given WT sequence, we launch the dead-end elimination (DEE) algorithm1 to optimize the mutant sequence base on the energy-like potentials (sequence potentials) inferred from the MSA. According to the Metropolis criterion, the design protocol will accept or reject a new mutant that may occur in the WT sequence, and the change is accepted with probability as follows, $$P=\min\{1, e^{-\Delta E/kT}\}$$
where \(\Delta E=E_{new}-E_{old}\) is the energy difference between the new and old mutant sequences, \(k\) is the Boltzmann constant, and \(T\) is the temperature.

Coupled mutations
Where are coupled residues located?
Based on the best-so-far sequence (designed), one can conduct a signle or multiple (coupled) mutants on the WT sequence, and the energy of the mutant sequence is computed for evaluating the conbination of mutants.
References:
- J. Desmet, M. D. Maeyer, I Lasters, Nature 356(6369):539, (1992)