Structural determinants of lipid specificity
Mitochondrial distribution and morphology protein 352020-10-13 O60200 · MDM35_YEAST
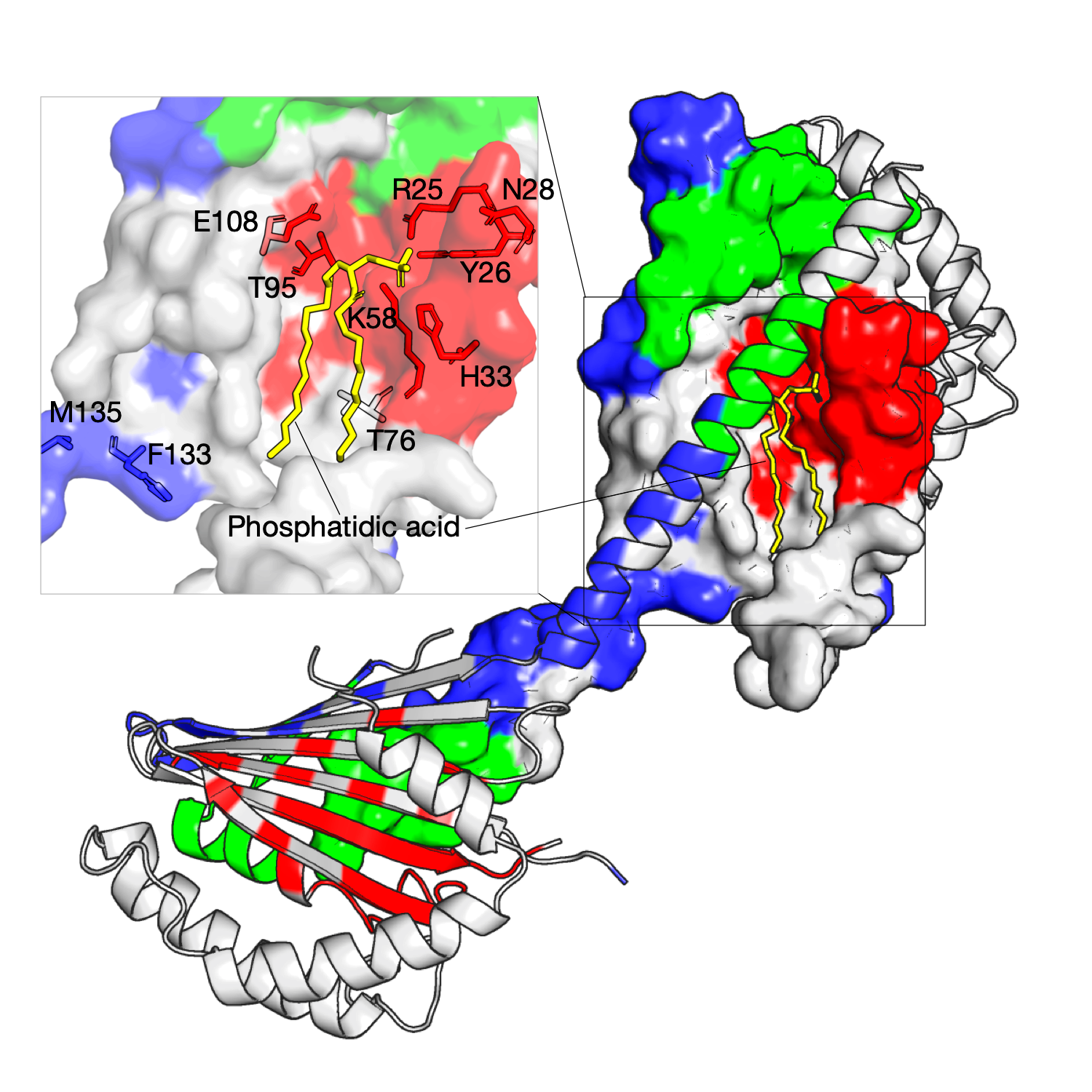
The Ups/PRELI lipid transfer proteins are widely conserved in eukaryotes and can be classified into two subfamilies by sequence similarity (Ups1/PRELID1 and Ups2/PRELID3. The Ups1–Mdm35 complex directly transfers phosphatidic acid (PA) between mitochondrial membranes allowing cardiolipin (CL) synthesis in the inner membrane. The gain-of-function Ups1 mutants from this approach can be categorised into two different functional regions: (a) residues facing to the inner substrate-binding cavity that tailor the cavity for selective accommodation of PA (K58, T95, E108) and (b) residues proximal to the Ω loop (K58, T76) and C-terminal α3 helix that likely have a role together in lipid extraction and capping an occupied binding cavity (F133, M135)1. As shown in the right figure, three residue communities are inferred from homologous sequences of the Ups1, and these residues in red are identified and mapped on the structure (PDB identifier: 4YTX, chain A, B, M, N), as shown, they are located at the tunnel-like binding cavity within the hydrophobic core.
Network of residues in the allosteric pocket govern Compound 1 inhibition and MKP5 selectivity
Dual specificity protein phosphatase 102020-09-22 Q9Y6W6 · DUS10_HUMAN
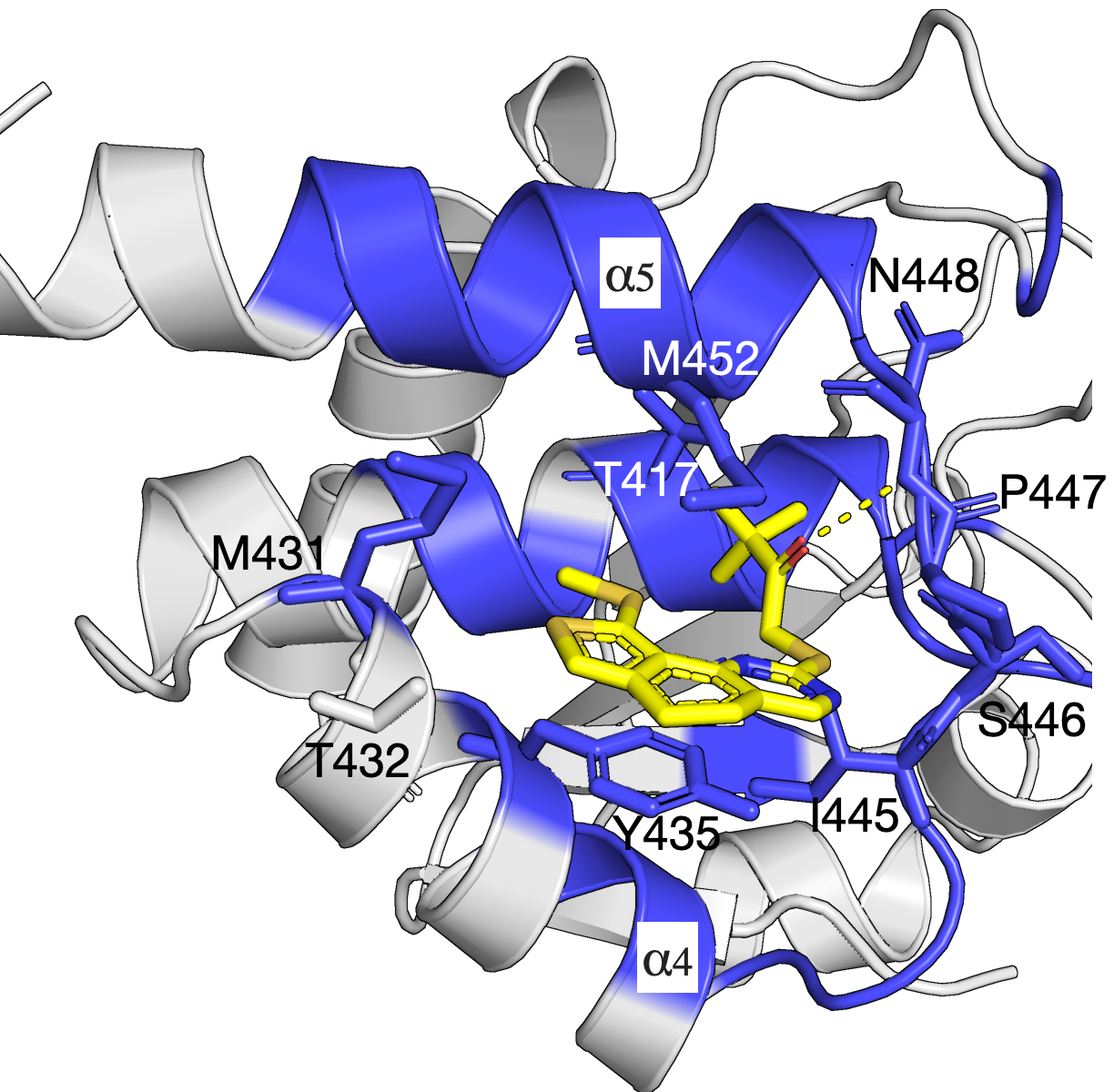
The mitogen-activated protein kinases (MAPKs) are a family of serine/threonine kinases that play indispensable roles in signal transduction pathways that control a plethora of physiological responses2. As shown in the left figure, an experimentally demonstrated allosteric binding pocket2 is identified by our SAEC method. In the analysis, only the homolog sequences are used to compute the evolutionary information and infer the binding pockect that agrees with that demonstrated in experiment. The identified network of residues is shown in blue and reveals key interactions between Compound 1 and MKP5-CD (PDB identifier: 6MC1).
Antagonistic activity on floral regulators FT and TFL1
Protein CENTRORADIALIS2020-09-18 Q41261 · CEN_ANTMA
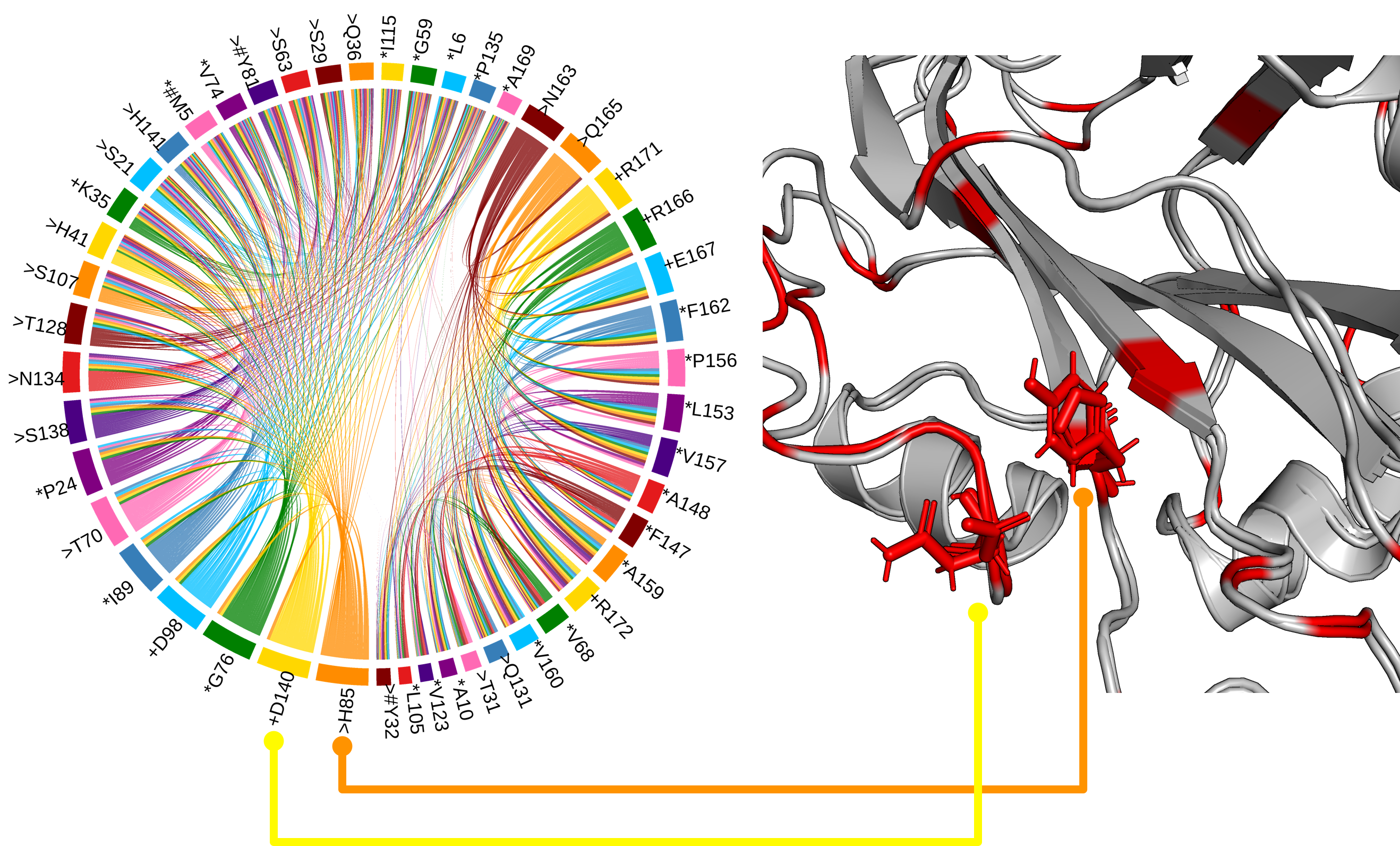
The His88 and Asp144 residues, which form a hydrogen bond in TFL1, but not FT, and which are likely the most critical residues for distinguishing FT and TFL1 activity3.
LetB tunnel is critical for function
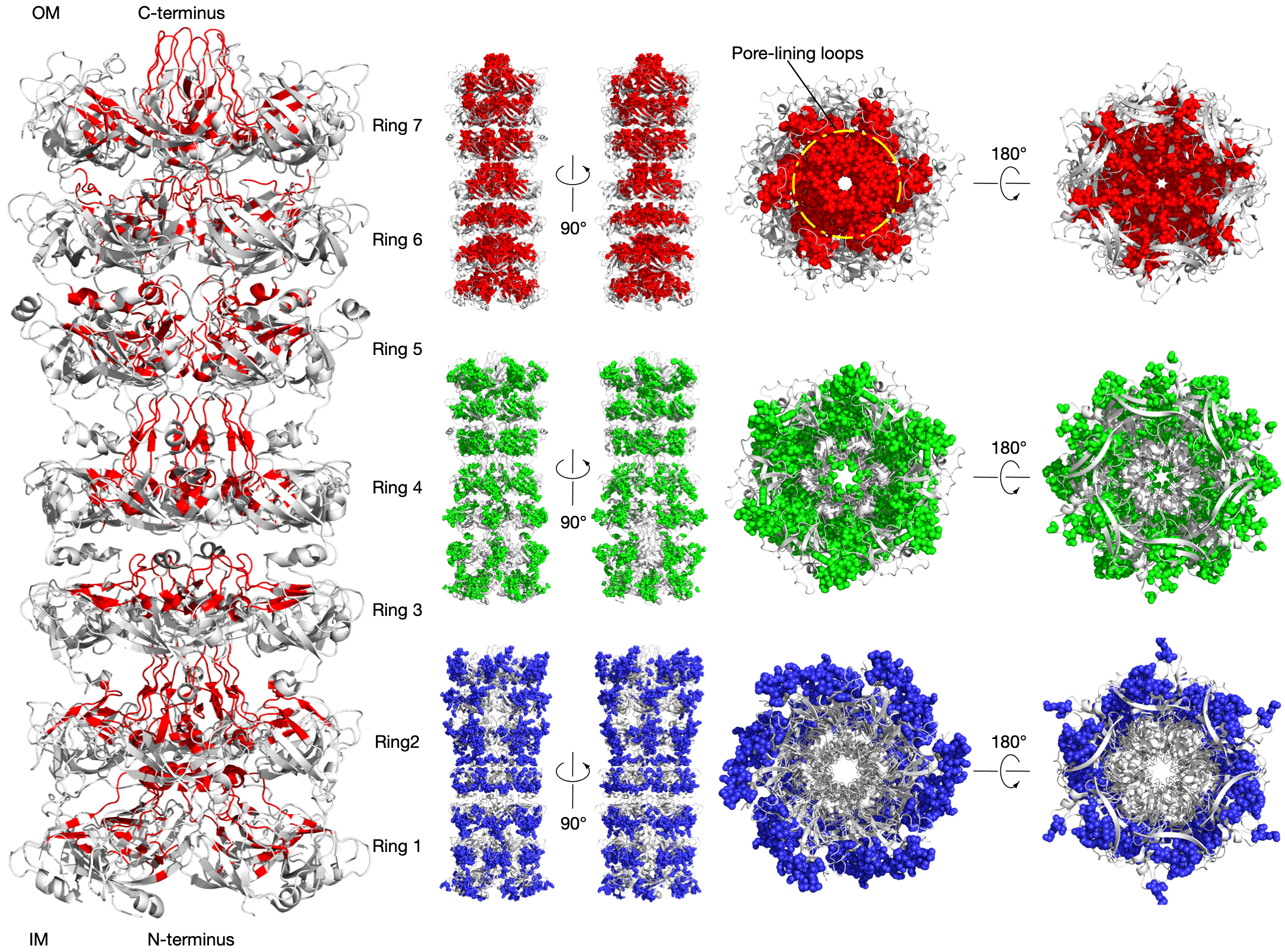

Intermembrane transport protein YebT2020-08-27 P76272 · YEBT_ECOLI
LetB forms a periplasm-spanning tunnel consisting of seven stacked rings and provides a hydrophobic pathway for the translocation of lipids across the periplasm4. Rings 1, 5, 6, and 7 are dynamic and adopt open and closed states. Our method has successfully identified communities in the LetB (PDB identifier: 6V0C) as shown in the figure (left). The pore-facing residues in red are inferred using AmoAi and contribute to the formation of the tunnel that mediates lipid transport in different states.
Bifurcated ligand-binding pocket in the ghrelin receptor
Growth hormone secretagogue receptor type 12020-08-25 Q92847 · GHSR_HUMAN
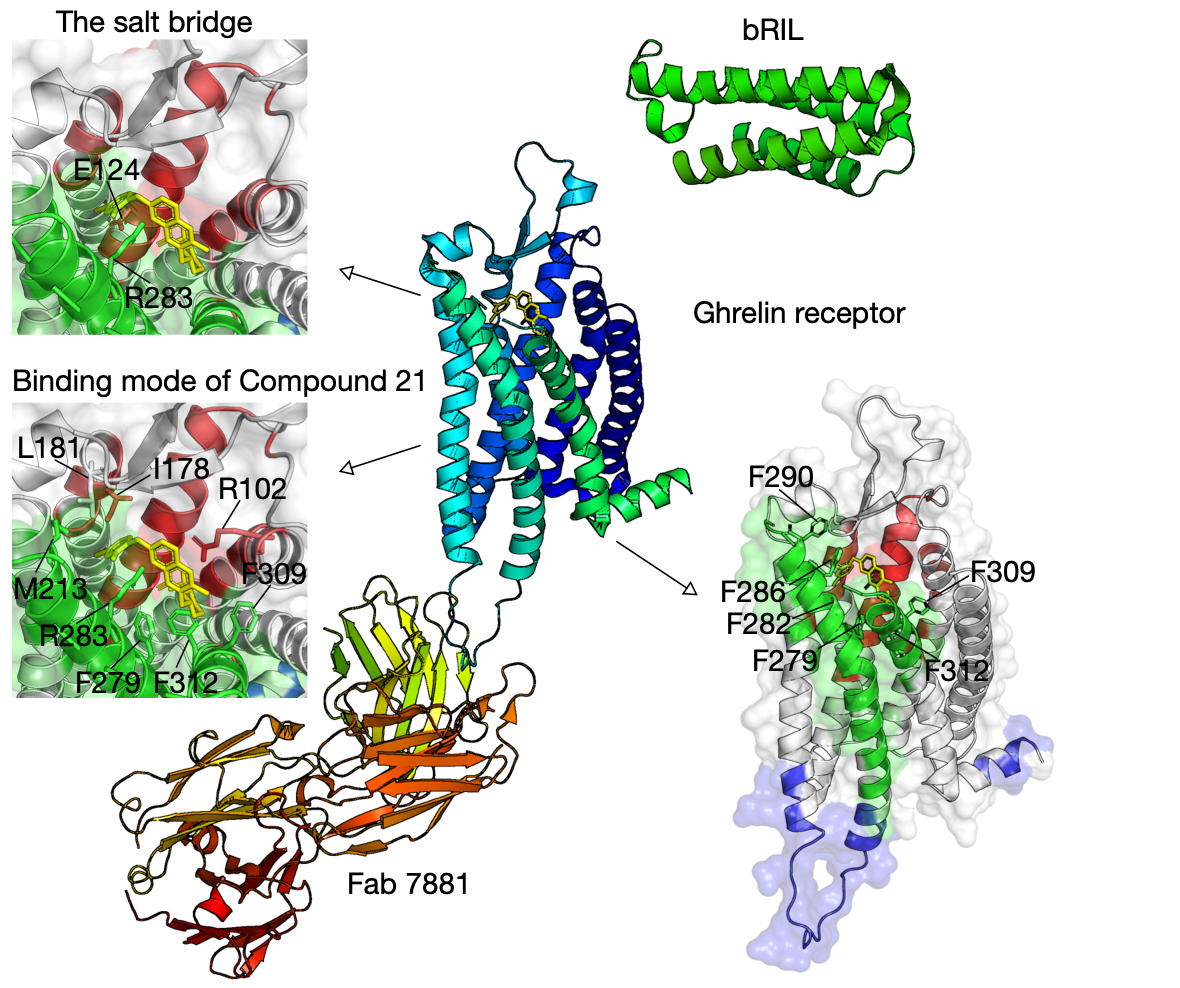
Ghrelin is a gastric peptide hormone with important physiological functions5. The unique feature of ghrelin is its Serine 3 acyl-modification, which is essential for ghrelin’s activity. Our method has successfully identified communities that cover the ligand-binding pocket of the ghrelin receptor (hGHSR PDB identifier: 6KO5), e.g., bifurcated by a salt bridge between E124 and R283.
The chorismate mutase
Bifunctional chorismate mutase/prephenate dehydratase2020-06-25 P0A9J8 · CMPDT_ECOLI
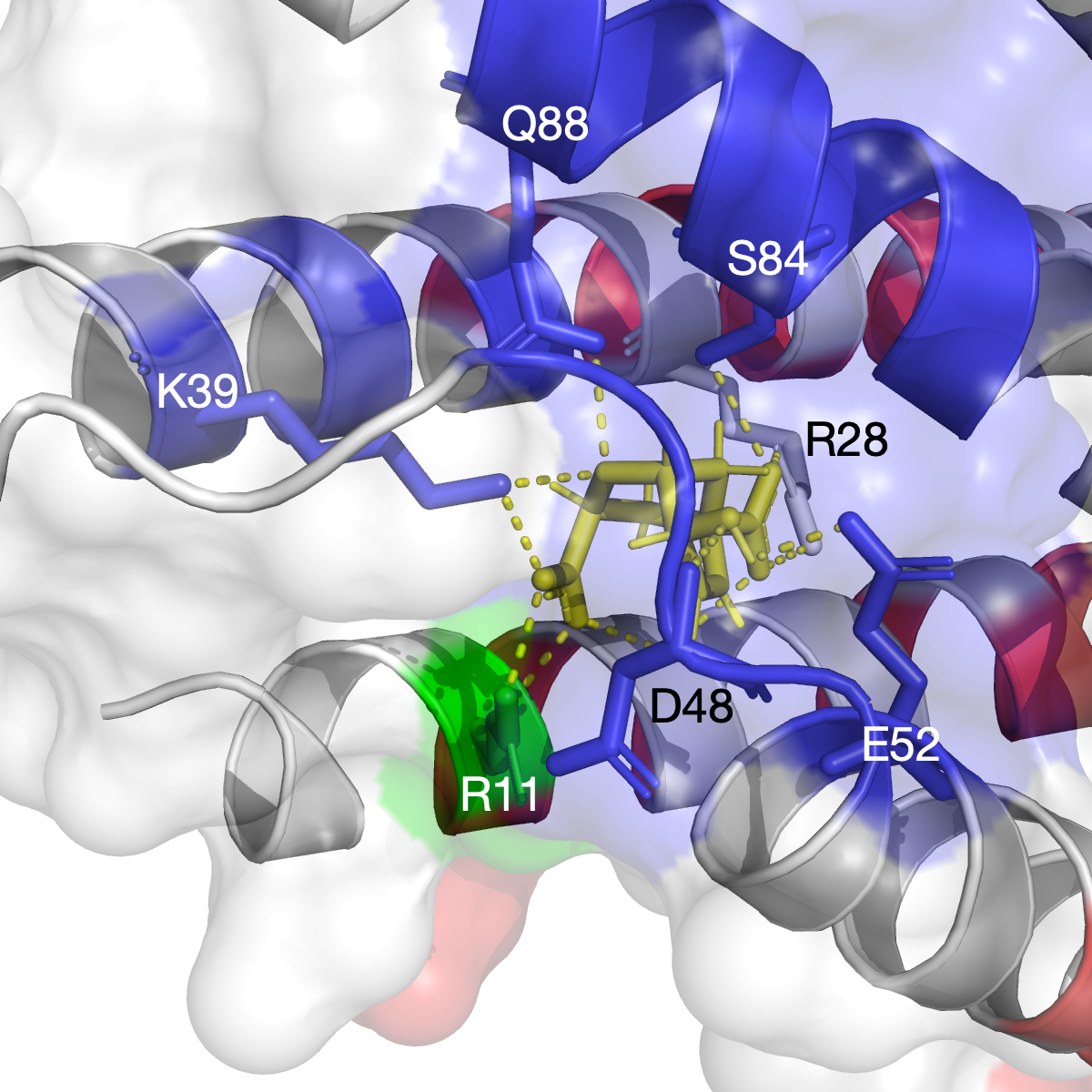
The chorismate mutase is an enzyme that is only found in fungi, bacteria, and higher plants, and it catalyzes the chemical reaction of converting chorismate to prephenate at the central branch-point of the shikimate pathway6. As a clssical system, it has been used to understand principles of catalysis and enzyme design. Our method has successfully identified communities in the chorismate mutase (PDB identifier: 1ECM) as shown in the figure (left). The residues that are functionally relevant to the catalytic activity are captured in the community (blue), and they are shown in stick.
The hinge domain of Smchd1
Chromosome partition protein Smc2022-01-07 Q9X0R4 · Q9X0R4_THEMA
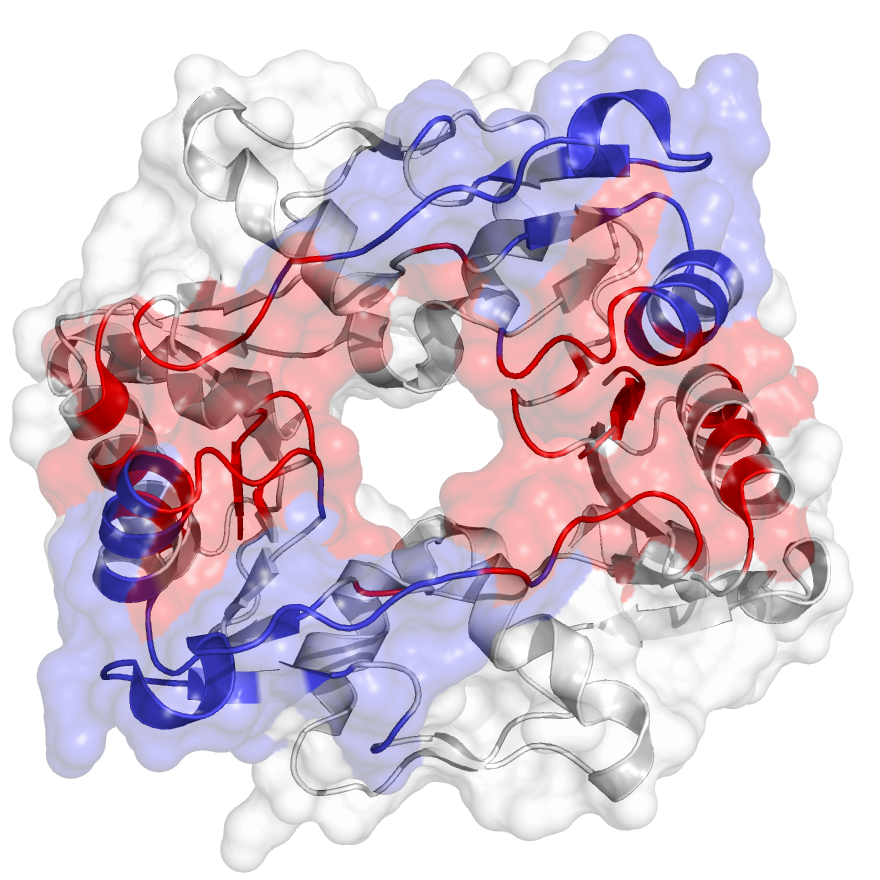
SMC flexible hinge domain containing 1 (SMCHD1) is a noncanonical member of the SMC protein family, and it has distinct features, e.g., an N-terminal gyrase, Hsp90, histidine kinase, and MutL (GHKL)–type ATPase domain and a hinge domain located at the C-terminus7. Disruption of SMCHD1 function is a potential factor that links to human disorders: the muscular dystrophy, facioscapulohumeral dystrophy, and the developmental disorder, Bosma arhinia micropthalmia syndrome. Our method has successfully identified communities that are functionally relevant to the canonical dimer interfaces of SMCHD1 hinge domain (PDB identifier: 1GXK).
References:
- Miliara, et al. Nature communications 10.1 (2019): 1-15.
- Gannam, et al. Science Signaling 13.646 (2020).
- Ahn, et al., The EMBO journal 25(3): 605-614, 2006.
- Georgia L., et al., Cell 181(3): 653-664, 2020.
- Shiimura, et al., Nature Communications 11(1): 1-9, 2020.
- Lee et al., JACS 117(12), 3627-8, 1995.
- Chen et al., Science Signaling 13(636), 2020.